Note: This documentation refers to CoCoGUI version 1.16
and may not be compatible with other versions.
Check the program itself for the actually supported options.
![]()
| back to the top |  |
![]()
CoCoNAD (for Continuous Time Closed Neuron Assembly Detection) is an algorithm for finding frequent parallel episodes or frequent (approximately) synchronous events in parallel point processes, for example, in parallel spike trains as they are recorded with multi-electrode devices in neurobiology. CoCoGUI is a Graphical User Interface for CoCoNAD, which is written in Java and accesses a C-based implementation of the CoCoNAD algorithm (JNICoCo, included in CoCoGUI). It also comprises the methods of Pattern Spectrum Filtering (PSF) and Pattern Set Reduction (PSR), with which the abundance of frequent patterns that are usually found in a given data set can be reduced to a smaller set of significant patterns.
If instead of a graphical user interface (as CoCoGUI provides it) a command line access to the CoCoNAD method as well as pattern spectrum filtering (PSF) and pattern set reduction (PSR) is desired, the Python scripts made available on the download page for the PyCoCo library (archive psf+psr.zip) are worth looking at. These scripts access the same C-based implementation of the CoCoNAD algorithm through a Python extension module written in C (the PyCoCo library), which also comprises the methods of Pattern Spectrum Filtering (PSF) and Pattern Set Reduction (PSR).
Of course, one may also write one's own Python scripts based on the PyCoCo library. Alternatively, there is an extension library for R, a statistical software package, namely CoCo4R.
General information about the theory underlying the CoCoNAD algorithm can be found in [Picado-Muiño and Borgelt 2013], while implementation aspects are discussed in [Borgelt and Picado-Muiño 2013]. The method of pattern spectrum filtering was introduced in [Picado-Muiño et al. 2013] (although for time-binned data, while CoCoNAD works on continuous data, which requires some adaptations) and was discussed in more detail as well as extended by methods for pattern set reduction in [Torre et al. 2013] (again for time-binned data, but the core ideas can be transferred to the continuous time operation of CoCoNAD). The method of pattern spectrum estimation was presented in [Borgelt and Picado-Muiño 2014]. It is recommended to study these papers before applying CoCoGUI to your own data. The following is only a very brief summary/review of some fundamental principles.
The core idea of CoCoNAD is to define a set of (approximately) synchronous events or a parallel episode in parallel (spike) trains or point processes as a group of events (action potentials or spikes in the neurobiology application) that occur with a user-specified maximum distance (in time) from each other. CoCoNAD then looks for frequent synchronous events or frequent parallel episodes, where the frequency or the support of a group of items (event types, or neurons in the neurobiological application) is measured by a maximum independent set approach: each synchronous occurrence of a set of items/events (spikes) of certain types (or: each parallel episode for the a certain set of items/event types, each set of approximately synchronous spikes from the same set of neurons) is an element of a family of sets. The size of a maximum independent set of this family, that is, of a selection of sets that have only empty pairwise intersections (independent set) and which comprises as many sets as possible (maximum independent set), is the support of the set of items/neurons. From a neurobiological point of view, this is a very plausible and intuitive support measure, since it ensures that no single event (spike) is counted more than once for the support of a given set of items/neurons. From a computer science/data analysis point of view, this support measure has the advantage that it is anti-monotone (or downward closed) and thus allows for effective and efficient pruning of the search for frequent synchronous events/parallel episodes.
The core idea underlying pattern spectrum filtering (PSF) is that among the abundance of frequent patterns found with CoCoNAD most are chance occurrences. That is, the majority of these patterns are such that they are likely to be observed even if there is no regularity in the (co-)occurrence of the events (that is, if the spike trains are independent or if events of different types occur independently). To remove these chance patterns and thus reduce the patterns to the (likely) significant ones, surrogate data sets are generated by randomizing and permuting the original data, with the aim of destroying any (regular) synchronous activity, but keeping as many other properties of the data as possible (like the number of events per train/spikes per neuron, local event/spike densities etc). These surrogate data sets implicitly represent the null hypothesis of independent point processes/(spike) trains. By generating and analyzing a sufficiently large number of surrogate data sets, a pattern spectrum is generated. A pattern spectrum maps pattern signatures, that is, pairs of the size of a pattern (its number of items/neurons) and its support (number of (co-)occurrences in the maximum independent set sense), to occurrence frequencies, that is, to the (average) number of patterns with such a signature that have been observed in the surrogate data sets. It is then assumed that any patterns found in the original data that possess signatures that occur also in at least one surrogate data set can safely be considered chance events (since a counterpart — same signature — occurred in presumably independent data) and thus can be removed from the patterns found in the original data.
As a faster alternative to generating a pattern spectrum by analyzing a large number of surrogate data sets, an estimation method is provided, which evaluates the characteristics of the data to produce an approximation of a pattern spectrum. Although this method yields only an approximation of an actual pattern spectrum, it has the advantage that it produces a pattern spectrum orders of magnitude faster and thus is the method of choice for larger data sets (more items/event types/neurons, longer recording periods) and for larger maximally allowed distances between events that are to be considered synchronous. Due to this speed advantage, estimating a pattern spectrum has been made the default operation mode. The pattern spectrum filtering, however, works in the same way in this case: only patterns with signatures that do not occur in the (estimated) pattern spectrum are kept.
The patterns that remain after pattern spectrum filtering are further reduced by analyzing subset and superset relationships between them, since chance coincidences of subsets of found patterns and chance coincidences of the whole set with items/neurons that are not part of an actual pattern can lead to smaller patterns with higher support and larger patterns with lower support, respectively, which are not significant themselves, but only induced by an actually significant pattern. The core principle of the pattern set reduction (PSR) step is to define a preference relation between patterns of which one is a subset of the other and then to keep only patterns to which no other pattern is preferred.
| back to the top |  |
![]()
The CoCoGUI program is distributed as a Java archive (cocogui.jar) and a library, which makes a C-based implementation of the CoCoNAD algorithm as well as an efficient, parallelized pattern spectrum generation available in Java through the Java Native Interface (JNI). Depending on the used operating system, this library comes either in the form of a so-called shared object (libJNICoCo.so, for GNU/Linux) or a so-called dynamic link library (JNICoCo.dll, for Microsoft Windows). Both the Java archive (system independent) and the library (system dependent — choose according to your system) are needed to run CoCoGUI. All of these files are available on the download page. (For Apple Macintosh/OSX, unfortunately, no precompiled libary can be provided, due to the lack of access to such a computer. However, it may work to compile the library from the sources, using the makefile for GNU/Linux. This has not been tested, though.)
On Microsoft Windows it should suffice to place the Java archive (cocogui.jar) and the dynamic link library (JNICoCo.dll) into the same directory and then to double-click the Java archive. (This presupposes, of course, that a sufficiently recent Java Runtime Environment (JRE) is installed, which is usually the case, because many programs, like web browsers, require it.) Alternatively, the batch file CoCoGUI.bat made available on the download page can be used to start the program. This batch file explicitly states that the current working directory should be searched by Java for the dynamic link library (JNICoCo.dll) and thus may avoid problems resulting from Java not being able to locate the library.
On GNU/Linux it is best to start the program with the shell script CoCoGUI.sh, which is made available on the download page. Directly double-clicking the Java archive (as for Microsoft Windows) may not work, because Java may not be able find the shared object (libJNICoCo.so) even if it resides in the same directory, because on a GNU/Linux system the current working directory is often not on the list of paths that Java searches for native libraries. The shell script explicitly adds the current directory to the Java library path and thus fixes this issue. Alternatively, the shared object may be copied to some directory that is on the Java library path. What directories are eligible may depend on the system configuration and thus cannot be stated generally.
On both Microsoft Windows and GNU/Linux, CoCoGUI may also be invoked from the command line with
java -jar -Djava.library.path="." cocogui.jar
Here the option -Djava.library.path="." adds the current working directory (denoted by ".") to the list of paths that are searched by Java for the library libJNICoCo.so (GNU/Linux) or JNICoCo.dll (Microsoft Windows). Note that the library can be placed into any other directory if the "." is replaced by the path to that directory. Note also that the batch file CoCoGUI.bat (for Microsoft Windows) and the shell script CoCoGUI.sh (for GNU/Linux) essentially only issue such a command, which may be adapted accordingly.
On successful invocation a window like this should appear (all screen shots shown here were made on a Ubuntu GNU/Linux system with a Gnome/Unity desktop environment):
| back to the top |  |
![]()
The main window of the CoCoGUI program features several tabs, on which the data files that the program reads and writes as well as several parameters for its operation can be specified. After the invocation of the program the second tab — that is, the "Files" tab — is in the foreground. To switch between the tabs, click on the tab headers on the left side of the window.

Below the tabs with their input fields a row with four buttons "Run", "Load", "Save" and "Close" is located. The "Close" button exits the program and closes the window. Alternatively, the program can be closed with the window manager button (in the screenshots on this page: the red circle with the diagonal cross in the top right of the window).
The "Run" button starts the execution of the core sequence of analysis operations of the CoCoGUI program, which include either estimating a pattern spectrum from the data characteristics or generating and analyzing surrogate data sets to obtain a pattern spectrum, analyzing the original data for frequent patterns, filtering the obtained patterns with the pattern spectrum (PSF), and finally reducing the remaining patterns to the relevant ones (PSR). With the example data file trains.txt, which is available on the download page, this can be tested directly (simply place this data file into the same directory from which CoCoGUI was started), without making any changes to the settings in any of the tabs. On a reasonably recent computer the analysis process, using pattern spectrum estimation, takes only about 0.5 seconds. If surrogate data sets are generated and analyzed to produce a pattern spectrum, the analysis should take between 20 and 60 seconds. In both cases, the analysis should finally produce 3 patterns. Note that it does not matter which tab is in the foreground when the "Run" button is pressed: the executed operations are always the same.
With the "Save" and "Load" buttons the current configuration of the graphical user interface (that is, the content and selections of the fields in the various tabs) can be saved to and loaded from a file. This may be useful if the same analysis is to be executed on several data sets on different runs of the CoCoGUI program, as it helps to avoid setting the same parameters each time.
If the CoCoGUI program is called (on a command line) with a file argument, that is, for example, with
java -jar -Djava.library.path="." cocogui.jar cocogui.cfg
or, using the start shell script (GNU/Linux), with
CoCoGUI.sh cocogui.cfg
or, using the batch file (Microsoft Windows), with
CoCoGUI.bat cocogui.cfg
the given file is interpreted as a configuration file that is to be loaded on start-up. That is, starting CoCoGUI in this way is equivalent to starting it without any file arguments, then pressing the "Load" button and selecting the configuration file given as an argument in the commands stated above.
| back to the top |  |
![]()
On the first tab — that is, the "Format" tab — the characters can be specified with which the data file is structured. The meaning of these characters and the choices for what a data record contains are explained in detail and with examples in the section on the data formats. The default settings are such that it should be possible to read common simple data formats without any changes.
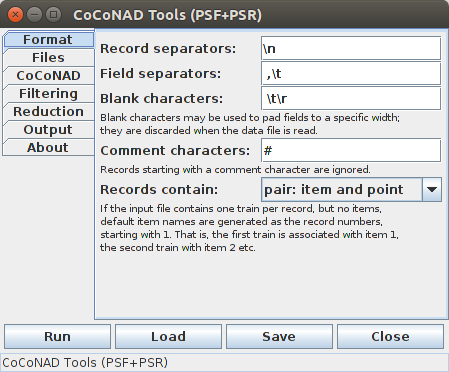
In the strings that are specified on this tab, standard ASCII espace sequences are recognized. For example, in the default setting, "\n" stands for the newline character, "\t" for the tabulator character and "\r" for the carriage return character. Furthermore, characters with special codes may be specified with "\000" or with "\x00", where the zeros have to be replaced by the octal code or the hexadecimal code of the character, respectively. The backslash character itself can be specified as "\\" or by making it the last character in a string.
| back to the top |  |
![]()
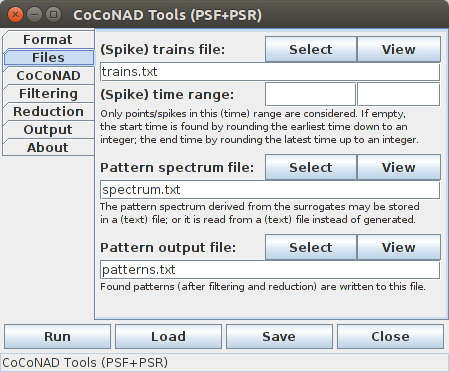
The second tab — that is, the "Files" tab — is the main tab of the CoCoGUI program. Here the input and output files are specified.

To test the operation of the CoCoGUI program, it is recommended to retrieve the example file trains.txt from the download page. If this file is placed into the same directory from which the CoCoGUI program was started, it will be used directly as the (spike) trains input data file. Pressing the "Run" button then executes the core sequence of analysis operations of the CoCoGUI program, which should end in a dialog box being displayed that informs about three patterns being found in the data.
To select a different data file, press the "Select" button next to the label "(Spike) trains file" in the first row of the "Files" tab and select the desired file in the file selector box that opens. Alternatively, the path name of the file may be typed directly into the text field under this label. This data file will always be read by the CoCoGUI program.
A (spike) time range may be specified, with which the input data is filtered. That is, only events/spikes with occurrence times in the specified time range are used in the analysis. The spike time range is also used to clamp the events/spikes in surrogate data sets that are generated to obtain a pattern spectrum. That is, no spikes/event times will be generated that lie outside of this time range. Technically, this is achieved by wrapping round spike/event times before the start of the range to its end and spike/event times after the end of the range to its start.
If the input fields for the (spike) time range are left empty, the spike time range is determined automatically from the data: the lower bound of the range is the earliest event/spike time rounded to the next lower integer (unless an integer itself), and the upper bound of the range is the latest event/spike time rounded to the next higher integer (unless an integer itself).
The pattern spectrum file is the file to which the generated pattern spectrum (that is, a mapping from pattern signatures to (average) occurrence counts) is written, which is obtained by generating and analyzing surrogate data sets. To select a name for this file, either type the path name into the text field under the label "Pattern spectrum file" or press the "Select" button to the right of it and then select the name of the file in the file selector box that opens. This file will be written by the CoCoGUI program, unless "read pattern spectrum from file" is selected as the surrogate data generation method on the "Filtering" tab. If this special method is chosen, the file specified here is read and its contents is used for the pattern spectrum filtering (PSF). Information about the data format of the pattern spectrum file can be found in the section on the data formats. Its format can be influenced to some degree on the "Output" tab.
The pattern output file is the file to which the found patterns (after pattern spectrum filtering and pattern set reduction) are written. This file is always written by the CoCoGUI program. Information about the data format of the pattern output file can be found in the section on the data formats. Its format can be influenced to some degree on the "Output" tab.
| back to the top |  |
![]()
On the third tab — that is, the "CoCoNAD" tab — various parameters for the CoCoNAD algorithm can be set, which influence what frequent patterns are found in the original and the surrogate data.
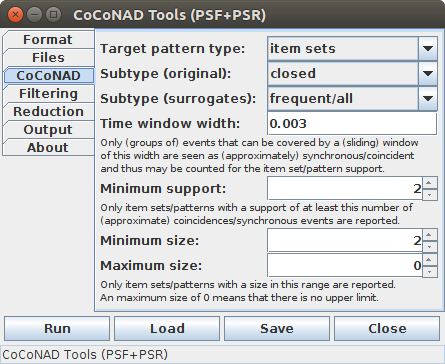
The CoCoNAD parameters start with a choice of the target pattern type, that is, whether item sets, partial item permutations (a.k.a. k-permutations or sequences without repetition) or item sequences (with and without repetition) should be found. With item sets the order in which the items occur in the sequence to analyze does not matter. With (partial) permutations, however, this order is respected: in all occurrences the items have to occur in the same order. As a consequence, (partial) permutations can yield multiple patterns that refer to the same set of items, since "a b c" and "b c a" are two different patterns as the order of the items differs. Finally, partial permutations differ from general sequences in that they do not allow for item repetition, that is, each item can occur at most once, while in general sequences the same item may occur multiple times in a found patterns.
Note, however, that neither for (partial) permutations nor for sequences the time distance between individual items is considered, but only whether the total of all items can be covered by a certain time window (same as for item sets).
The main target type (item sets, (partial) permutations or sequences) is the same for both the original data and the surrogate data. However, the target pattern subtype may differ for these data sets. There are three options: "frequent/all", "closed" and "maximal". A pattern is called frequent if it has a support (in the maximum independent set sense, see above) that is at least the user-specified minimum (to be specified further down on this tab). A pattern is called closed if there is no super-pattern that has the same support (or, in other words, all super-patterns have a lower support — note that super-patterns cannot have a higher support, because support is anti-monotone). A pattern is called maximal if there is no super-pattern that is frequent. Closed and maximal patterns are also required to be frequent, but are a restriction of all frequent patterns (explaining the "all" in the first option), as defined by the conditions stated above.
It is recommended to use closed patterns for analyzing the original data, in order to avoid subpatterns with the same support (even though (most) of these should be removed by pattern set reduction) and all frequent or also closed frequent patterns for the surrogate data sets. Although it may sound more plausible to use closed frequent patterns in both cases for consistency, the pattern spectrum filtering that is carried out is the same, because the decision border (the border between occurring and non-occurring pattern signatures, which governs the pattern spectrum filtering) is the same in both cases, because all closed frequent patterns are obviously also frequent patterns (by definition) and the decision border is determined only by the closed frequent patterns. This makes finding all frequent patterns preferable for the surrogate data sets, because it is usually somewhat faster and needs less memory, while producing the same final results. Note that this selection for the analysis of surrogate data sets is ignored if a pattern spectrum is estimated from data characteristics, without ever producing any surrogate data sets ("estimate pattern spectrum" chosen on the "Filtering" tab). An estimated pattern spectrum always refers to all frequent patterns.
How far the events/spike times of an (approximately) synchronous event or a parallel episode may be apart is specified as a (time) window width (or maximum time distance). It has to be stated in the (time) unit that is used in the (spike) train file. For an application in spike train analysis, where the time unit is usually seconds, three milliseconds (or 0.003 seconds) appears to be a reasonable choice. This value should be chosen with care, since large values can lead to prohibitively long processing times, especially if actual surrogate data sets are to be analyzed.
What patterns count as frequent is specified with the minimum support. It states the minimum size of a maximum independent set of (approximately) synchronous events or parallel episodes that has to be reached in order to render a group of items/neurons a frequent pattern. Only frequent patterns are reported or generated from the original and surrogate data sets, respectively. It is recommended to use a minimum support of at least two and to increase this value if the analysis process takes too long.
The size of the patterns to be found can be restricted by a minimum and a maximum. It is recommended to use a minimum of at least two items (event types, neurons) and not to restrict the maximum size (in order to avoid horizon effects, which can occur if only closed or only maximal frequent patterns are to be found). The latter is achieved by specifying a maximum size of zero, which is the default setting. If the maximum size is less than the minimum (but greater than zero), it is (internally) automatically replaced by the minimum size for consistency.
| back to the top |  |
![]()
On the fourth tab — that is, the "Filtering" tab — the parameters for the pattern spectrum filtering (PSF) can be specified. The content of this tab is context dependent. If for the surrogate data generation method any but the "pattern spectrum estimation" entry is selected, the tab looks like this (only for "dithered blocked event/spike permutations" the "Block size" input is enabled, though):
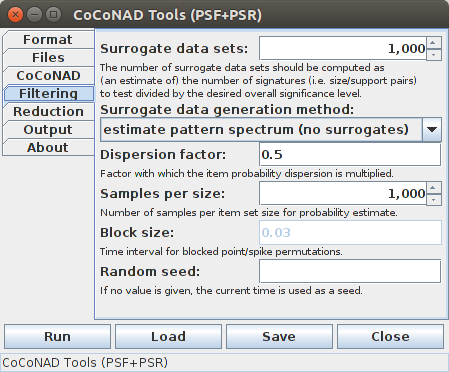
If, however, "pattern spectrum estimation" is chosen instead of actually generating and analyzing surrogate data sets, the tab looks like this:
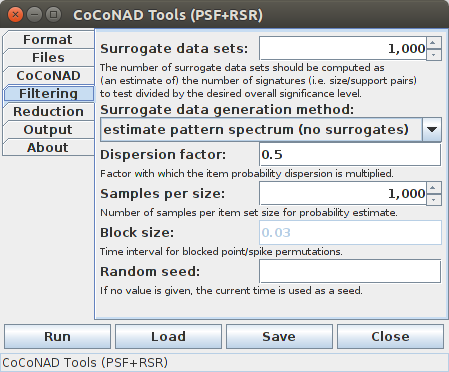
The PSF parameters start with the number of surrogate data sets to generate, or the equivalent number of such surrogate data sets if a pattern spectrum is to be estimated (which is used to round and threshold the estimated counts). According to [Picado-Muiño et al. 2013] this number should be computed as an estimate of the number of pattern signatures that occur in the original data and one is actually willing to consider as significant, divided by the desired significance level for the patterns (thus implementing a simple Bonferroni correction for multiple testing). For example, if there are 30 pattern signatures that one is actually willing to consider as possible patterns and the desired significance level is 1% = 0.01, then 30/0.01 = 3,000 surrogate data sets should be generated. However, for most practical purposes, 1,000 data sets are usually sufficient, since the decision border does not change much with the number of surrogate data sets, provided it is not too small. The procedure has also been found to be fairly conservative.
For generating surrogate data sets, six different methods are available, plus two special options for not performing pattern spectrum filtering ("none (no pattern spectrum filtering") and keeping the original data unchanged ("identity (keep original data"). Since generating multiple surrogate data sets with an identity mapping is obviously useless (as the exact same set of patterns will be generated every time), the number of surrogate data sets is ignored for this option, and only one surrogate data set is generated (which, due to the identity mapping, is identical to the original data). This option mainly serves the purpose to allow a user to generate and store the pattern spectrum of the original data. Furthermore, the pattern spectrum to be used for filtering can be read from a file ("read pattern spectrum from file", where the file name has to be specified on the "Files" tab).
Most of the actual surrogate data generation methods require a probability density function, from which random numbers are sampled, and a parameter that governs the width of this density function ("Sigma parameter"). The density function can be "rectangular" (or uniform), "triangular" or "Gaussian/normal". For the first two choices the sigma parameter states half the base width of the rectangular or triangular density function, in the same (time) units that are used in the (spike) train data file (and in which the time window width on the "CoCoNAD" tab is stated as well). For a Gaussian/normal density function, the sigma parameter states the standard deviation (which explains the name "sigma parameter", since in statistics the standard deviation is usually denoted by σ — the Greek character sigma).
The surrogate data generation methods are:
Note that the density function type and sigma parameter are ignored for the first surrogate data generation method, because this method always samples from a uniform distribution on the (time) range of the events/spikes.
Note also that the last three methods can, in principle, be applied with a sigma parameter of zero (that is, the chosen events/spikes are not displaced/dithered), while the second and third method require a non-zero sigma parameter to actually modify the data. The reason is that the last three methods not only displace/dither events/spikes in a train, but also permute events/spikes between trains. This has the advantage that possibly present synchronous activity is more thoroughly destroyed. However, the fourth and the fifth method have the disadvantage that rate profiles of individual trains may also be changed (only the overall rate profile — sum over all trains — is preserved). Only the last method preserves (up to the precision of the block size) the rate profiles of individual trains.
The execution of all surrogate generation methods is parallelized internally. That is, as many surrogate data sets are generated and analyzed in parallel as the computer on which CoCoGUI is running reports as available processor cores. This makes the pattern spectrum generation much faster, but can render the computer unresponsive and slow as long as the analysis is running, since all available processing power of the machine is utilized.
In order to make the surrogate data generation reproducible, a seed for the pseudo-random number generator can be specified. An empty seed or a seed value of zero mean that the current time (as a Unix time stamp) is used as a seed (and thus renders the surrogate data generation essentially irreproducible). Note, however, that reproducible results can only be obtained on computers that offer the same number of processor cores. A different number of processor cores usually means that a different set of surrogate data sets is generated, due to the different split into parallel execution threads.
As an alternative to generating and analyzing surrogate data sets, a pattern spectrum may be obtained by estimating it from the characteristics of the (spike) trains:
The pattern spectrum estimation essentially assesses the expected counts for the pattern signatures for each pattern size separately. For a given pattern size, the number of "slots" in the data are counted (that is, the number of event groups of the given size that fall within the specified window width/are no farther apart than this maximum distance) that can hold a pattern of this size. Then a distribution over the coincidences/support values is estimated as an average of Poisson distributions, where the distribution parameter is computed from the number of slots and estimates of the probability of specific item/event type sets. This probability is computed from the occurrence probabilities of the individual items/event types. However, it turns out that using the item/event type probability as it can be derived from the data leads to an overestimate of the expected number of coincidences. As a heuristic correction for this, the dispersion of the item probabilities is multiplied by a factor (less than 1) in order to make the probabilities more similar than they actually are. Empirically its was found that a factor around 0.4 to 0.5 leads to good results. This factor can be entered after the label "Dispersion factor". Furthermore, since different items/events occur with different probabilities and it is (due to a combinatorial explosion) impossible to enumerate all possibilities, samples of concrete item/event type sets are drawn. How many samples are to be drawn per item set size (and thus how many Poisson distributions are to be averaged for an estimate of the distribution over the coincidences/support values) can be specified after the label "Samples per size". Since the sample drawing is also a random process, which one may want to make reproducible, a seed for the random number generator may be specified (see the explanations as they were given above for the surrogate data generation). Note that the number of surrogates is also used for the pattern spectrum estimation, namely as an equivalent number which determines how the occurrence counts for the signatures are rounded and thresholded.
After a pattern spectrum has been obtained, either by analyzing surrogate data sets or by estimation, it is written to the pattern spectrum file that is specified on the "Files" tab and the patterns found in the original data are filtered with it. That is, only patterns with signatures that do not occur in the surrogate data sets/in the (estimated) pattern spectrum are kept, where a pattern signature is a pair of pattern size (number of items/neurons) and pattern support (occurrence frequency/number of coincidences in a maximum independent set sense). All other patterns are discarded.
| back to the top |  |
![]()
On the fifth tab — that is, the "Reduction" tab — the parameters for the pattern set reduction (PSR) can be specified. There is (currently) only one, namely the pattern set reduction method:
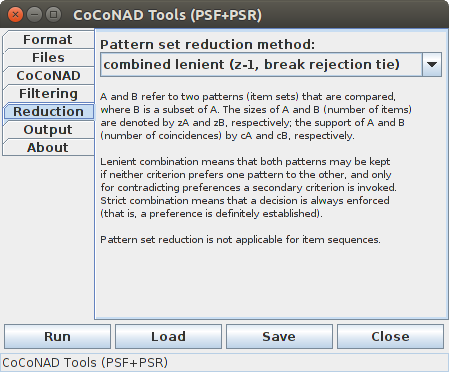
Pattern set reduction (PSR) further reduces the patterns that remain after pattern spectrum filtering (PSF). It analyzes subset and superset relationships between the found patterns. The reason for this step is that due to chance coincidences of subsets of items/neurons of an actually significant pattern or a few chance coincidences with additional items/neurons outside of the actually significant patterns, the filtered pattern set may still contain non-significant induced patterns. These patterns are removed based on the following rationale: between pairs of patterns, one of which is a subset of the other, a (possibly incomplete) preference relation is defined. Based on this preference relation, only those patterns are kept to which no other pattern (subset or superset pattern) is preferred.
In the following description of the pattern set reduction methods (or rather the preference relations underlying them), A and B refer to two patterns (sets of items/neurons) with B ⊆ A (that is, B is a subset — or subpattern — of A), z refers to the size of the pattern (that is, the number of items/neurons contained in it), and c refers to the support of the pattern (that is, the number of coincidences/synchronous events/parallel episodes in a maximum independent set sense exhibited by the pattern).
The pattern set reduction methods/preference relations are:
After the set of patterns has been reduced with pattern set reduction (PSR), the final set of patterns is written to the pattern output file specified on the "Files" tab.
| back to the top |  |
![]()
On the fifth tab — that is, the "Output" tab — parameters can be specified that influence the format of the files that are written by the CoCoGUI program.
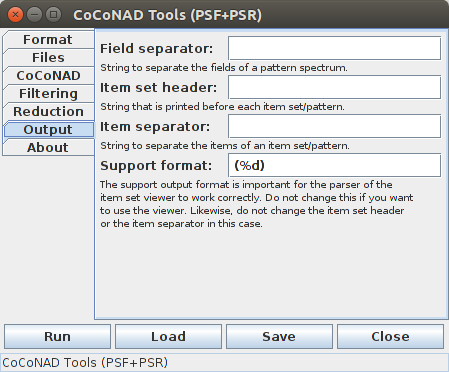
The parameters on this page should be handled with extreme care, because many choices can make it impossible to read the files into the viewers that are described below. Therefore it is recommended to leave these parameters at their default settings, unless you really know what you are doing.
The "Field separator" states the string that is printed between the fields of a pattern spectrum (a general description can be found in the section on the data format). In principle, this can be any string, but only a space (the default), a tabulator, or a comma ensure that it is possible to properly read a pattern spectrum into the viewer that is described below.
The "Item set header" specifies a (fixed) string that is to be printed before each item set/pattern in the pattern output file. By default, this string is empty. Using a non-empty string may lead to errors or strange effects when reading a pattern set into the viewer that is described below.
The "Item separator" states the string that is printed between the items of a found pattern. In principle, this can be any string, but only a space (the default), a tabulator, or a comma ensure that it is possible to properly read a pattern set into the viewer that is described below.
The "Support format" specifies how the support information should be formatted after the items of a pattern. The "%d" states that the support should be printed as a decimal integer number (standard C and Java format string). The support is surrounded by parentheses in order to distinguish it from the items of the pattern (which may be numbers as well) and preceded by a separator character (a space by default), which may differ from the item separator. Apart from adding additional spaces around the parenthesized expression, essentially all changes to this format string will render it impossible to read the pattern set into the viewer that is described below.
| back to the top |  |
![]()
The last tab — that is, the "About" tab — provides version and license information about the CoCoGUI program.
| back to the top |  |
![]()
The CoCoGUI program includes viewers for all file types that it reads or writes, which can be invoked with the "View" buttons on the "Files" tab (next to the "Select" buttons, with which the files can be selected).

The "View" button next to the label "(Spike) trains file" invokes a viewer for parallel spike trains/point processes in the form of a dot display, the "View" button next to the label "Pattern spectrum file" invokes a table viewer for a pattern spectrum that has been generated by analyzing surrogate data sets. This viewer allows to display the table as a 3D bar chart or a 3D scatter plot. Finally, the "View" button next to the label "Pattern output file" opens a tabular viewer for the found patterns. These viewers are described in more detail in the following sections.
| back to the top |  |
![]()
The "View" button next to the label "(Spike) trains file" on the "Files" tab opens a simple dot display for parallel spike trains/point processes.
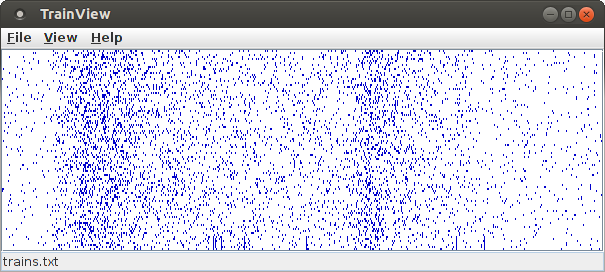
This dot display shows the example data file trains.txt that is available on the download page. Each row of this dot display, which is two pixels high by default, corresponds to an item/neuron. The horizontal axis corresponds to the time domain and is scaled with 200 pixels per (time) unit (as it is used in the data input file). The default size of the window is 600 times 200 pixels and thus exactly captures the data file, which contains 100 items/neurons recorded for 3 seconds.
The items/neurons will be shown ordered from bottom to top. That is, the events/spikes of the item/neuron with the smallest name/identifier will occupy the bottom row, the events/spikes of the item/neuron with the greatest name/identifier will occupy the top row. If all item names/identifiers are numbers, the items/neurons are ordered by the numeric value of their names/identifiers, otherwise they are ordered lexicographically.
Note that in this display one set of items/neurons that exhibits synchronous activity is immediately visible, namely the set consisting of the items/neurons associated with the bottom seven rows. In these rows seven vertical lines are visible, which correspond to the seven synchronous events/parallel episodes exhibited by these seven items/neurons. However, this pattern is not the only one present in this data set. There are two other sets of items/neurons (with five and nine items/neurons, respectively) that exhibit synchronous activity. These patterns are not directly visible, because the involved items/neurons are not located side by side (that is, in adjacent rows) in this display. This demonstrates that a dot display is, in general, not sufficient to discover (all) patterns that are present in the data, but that more sophisticated methods are necessary that are able to efficiently check the vast number of subsets that could potentially form patterns.
The layout of the dot display can be influenced with the dialog box that can be opened with the menu item "View > Set Layout ...".
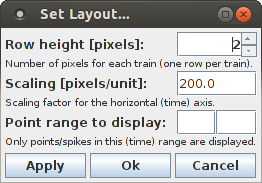
This dialog box allows to change the height of each row in pixels, the scaling for the horizontal (time) axis (in pixels per unit) and the event/spike time range to display. If the event/spike time range is left empty, it is automatically determined from the data: the lower bound of the range is the earliest event/spike time rounded to the next lower integer (unless an integer itself), and the upper bound of the range is the latest event/spike time rounded to the next higher integer (unless an integer itself). The dialog box has three buttons: "Apply" applies the settings in the dialog box to the bar chart and keeps the dialog box open to make additional changes. "Ok" applies the settings and closes the dialog box. "Cancel" merely closes the dialog box, but does not apply the settings to the bar chart.
In addition, it is possible to change the color of the dots in the dot display with a dialog box that is opened with "View > Set Color ...". This dialog box has the standard form of a Java color selection dialog and allows to choose any possible color for the dots in the display.
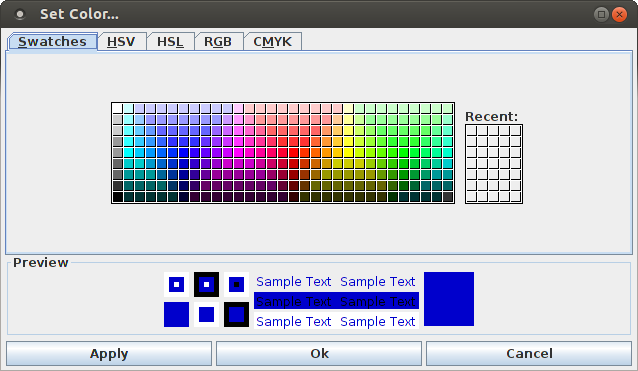
Note the different tabs of this dialog box that correspond to different color models/color selection methods. Note also that this dialog box has the same three buttons with the same functions as the layout dialog box.
| back to the top |  |
![]()
The "View" button next to the label "Pattern spectrum file" on the "Files" tab opens a simple table viewer for the elements of a pattern spectrum. Note that a pattern spectrum is only available after the analysis process has been executed by pressing the "Run" button of the main CoCoGUI window and waiting for completion. Note also that due to the random nature of the surrogate data generation, the pattern spectrum may differ from the one shown in the following screen shots.
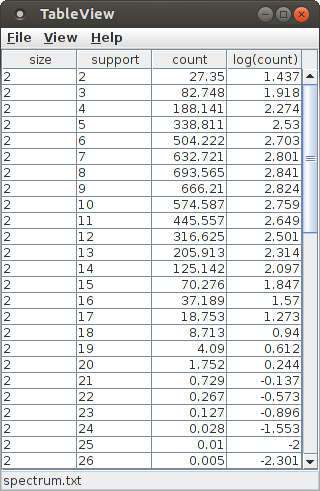
A pattern spectrum maps pattern signatures to their (average) occurrence counts among patterns found in the surrogate data sets, where a pattern signature is a pair consisting of the pattern size (number of items/neurons) and pattern support (number of coincidences). For convenience, especially w.r.t. a visualization as a 3D bar chart or 3D scatter plot the logarithm of the occurrence count is added to the columns of the table.
The pattern spectrum can be visualized as a 3D bar chart by selecting "View > BarChart ..." from the menu.
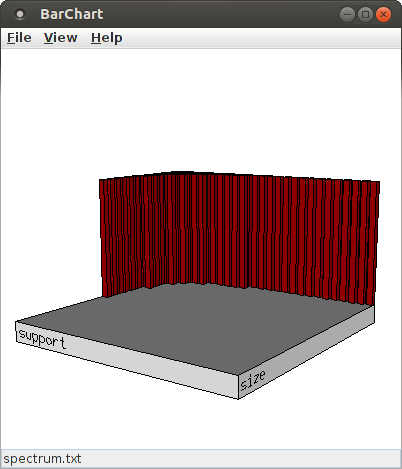
The initial bar chart view that is opened is a simple histogram of pattern size and support. That is, the bars merely indicate, which combinations of pattern size and support occurred among the patterns found in the surrogate data sets. In the initial view, the size is on the horizontal axis, the support on the axis pointing into the drawing plane. This view can be changed in several ways using the mouse. The most direct way is to press the left mouse button at any point in the diagram and keep it pressed while moving the mouse. This has an effect as if a point on a sphere that is centered in the middle of the base plate is grabbed and the sphere turned with it. By pressing the middle mouse button and keeping it pressed while moving the mouse, the chart can be rotated around an axis that points into the drawing plane and is centered in the window. By pressing the right mouse button and keeping it pressed while moving the mouse, one can zoom into or out of the chart. Pressing and holding the shift key or the control key on the keyboard modifies the behavior of the mouse buttons. With a pressed shift key, pressing the left mouse button and keeping it pressed while moving the mouse rotates the chart around a horizontal axis and with the right mouse button around a vertical axis, while the middle mouse button still rotates around an axis pointing into the drawing plane. With a pressed control key, the location of the chart in the window can be changed. With the left mouse button, the chart can be moved in both direction, with the middle mouse button only in horizontal direction and with the right mouse button only in vertical direction.
To change what is displayed in the chart, an attribute selection dialog box can be opened with "View > Set Attributes ...".
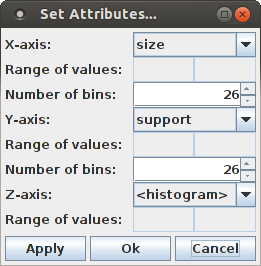
Since for displaying a pattern spectrum as a bar chart it is best to choose the pattern size and pattern support for the base axes, it is not recommended to change the selection for the X-axis or the Y-axis. Only changing the number of bins to be displayed can be useful (see below). The most illustrative display is achieved by selecting the logarithm of the (average) occurrence counts of a pattern signature in the surrogate data for the Z-axis:
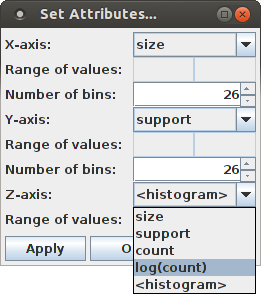
Note that "<histogram>" in this list refers to the initial bar chart view, in which a bar only indicates whether a pattern signature occurs among the patterns found in the surrogate data sets or not.
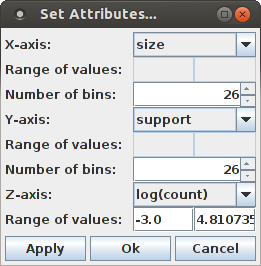
Selecting "log(count)" from the list of attributes for the Z-axis chooses the (average) occurrence counts of a pattern signature for the Z-axis. Then pressing "Apply" (apply the settings, but keep the dialog box open for further changes) or "Ok" (apply the settings and close the dialog box) yields the following display (while "Cancel" merely closes the dialog box without making any changes to the bar chart):
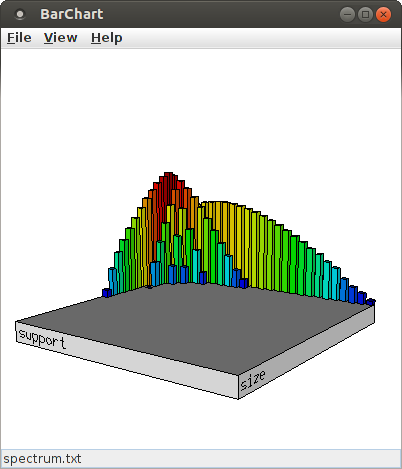
Since the number of bars can make it difficult to explore details around the relevant signatures, it can be useful to reduce the number of bins for the X- and Y-axis. For this, change the content of the spinners next to "Number of bins", either using the small up and down arrows or by typing a new value into the input field. For example, with 14 bins for both the size and the support direction (and the scaling for the bar height increased to 160%, see below), the bar chart looks like this:
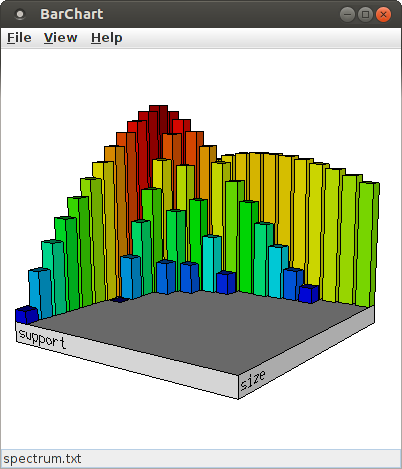
The layout of the bar chart can be changed by opening the layout dialog box with "View > Set Layout ...".
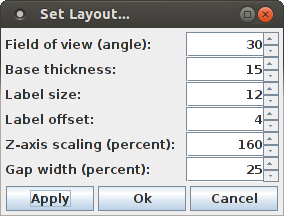
The layout dialog box allows to change the field of view of the projection, which is stated as an angle (namely the angle at which the viewing frustum opens at the eye point). The 3D projection is generally a central projection, which is the closer to a parallel projection, the smaller the angle that defines the field of view.
The base thickness is the height of the base plate of the bar chart, stated in pixels for a parallel projection (as a consequence of the central projection, the base plate height may be greater or smaller than 15 pixels). The label size specifies the size of the labels that are written on the sides of the base plate, again in pixels in a parallel projection. The label size should not be larger than the base thickness. The label offset states how many pixels the base line for the label text is above the bottom edges of the base plate. With the Z-axis scaling the height of the bars can be influenced. For the bar chart with 14 bars for both the pattern size and the pattern support direction shown above, the Z-axis scaling was increased to 160% (the default is 100%). Finally, the gap width controls how wide the gap between the bars of the bar chart is, stated as a percentage of a grid cell. For example, with the default 25%, the gap is one quarter of a grid cell wide, while the bar fills the remaining three quarters.
The scheme used to color the bars, which by default is a rainbow scale that encodes the bar height, can be modified with the color dialog box that opens if "View > Set Color ..." is selected.
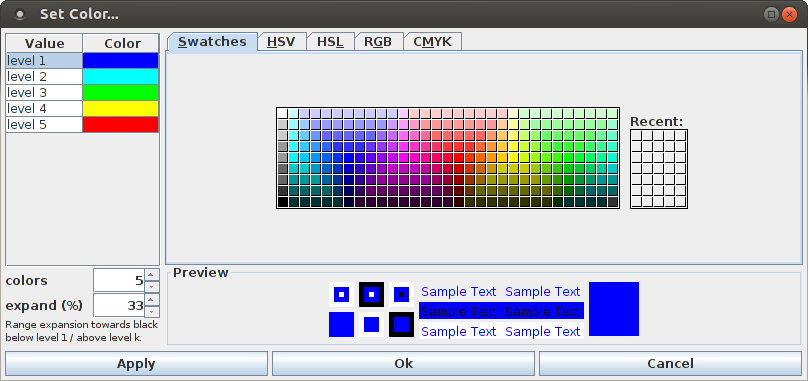
The right part of the color dialog box is a standard Java color selector, as it is also used for selecting the color of the dots in the dot display used to visualize parallel point processes/spike trains (see above). The left part of the dialog box defines a list of colors that is used to specify the color range that is used to encode the bar height. The default is a list of five color levels, ranging from blue to red, with turquoise, green and yellow in the middle. The number of colors can be changed with the "colors" spinner below the color list, for example, it can be reduced to two colors to obtain a simple heat map scale. The second spinner, labeled "expand (%)" specifies by how much the color range is extended towards black below the color for level 1 and above the color for level k, where the percentage refers to the step between two colors (100%). An expansion factor of one third (33%) usually yields a visually pleasing color scale.
As an alternative to a bar chart, a pattern spectrum may also be visualized as a 3D scatter plot, which can be selected with "View > ScatterPlot ..." in the pattern spectrum table viewer.
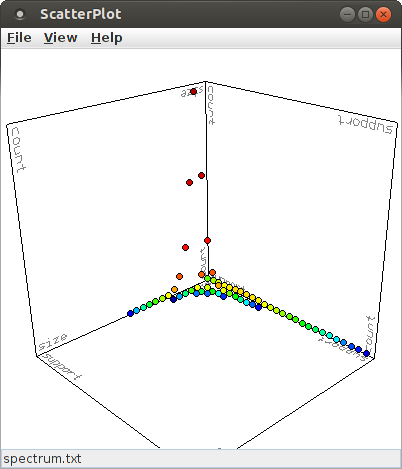
In the default view, the X- and Y-axis refer to the pattern size (number of items/neurons) and the pattern support, respectively. The (average) number of occurrences of a pattern signature in the surrogate data sets is used for the Z-axis and the color of the points is computed from the logarithm of this number of occurrences. This view can be changed in several ways using the mouse. The most direct way is to press the left mouse button at any point in the diagram and keep it pressed while moving the mouse. This has an effect as if a point on a sphere that is centered in the middle of the diagram cube is grabbed and the sphere turned with it. By pressing the middle mouse button and keeping it pressed while moving the mouse, the plot can be rotated around an axis that points into the drawing plane and is centered in the window. By pressing the right mouse button and keeping it pressed while moving the mouse, one can zoom into or out of the plot. Pressing and holding the shift key or the control key on the keyboard modifies the behavior of the mouse buttons. With a pressed shift key, pressing the left mouse button and keeping it pressed while moving the mouse rotates the plot around a horizontal axis and with the right mouse button around a vertical axis, while the middle mouse button still rotates around an axis pointing into the drawing plane. With a pressed control key, the location of the chart in the window can be changed. With the left mouse button, the chart can be moved in both direction, with the middle mouse button only in horizontal direction and with the right mouse button only in vertical direction.
The selection of attributes that is used for the scatter plot can be changed by opening the attribute selection dialog box with "View > Set Attributes ...".
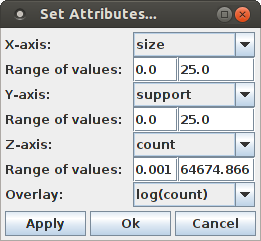
A more illustrative view is obtained by choosing the logarithm of the (average) occurrence count for the Z-axis, which yields this diagram:
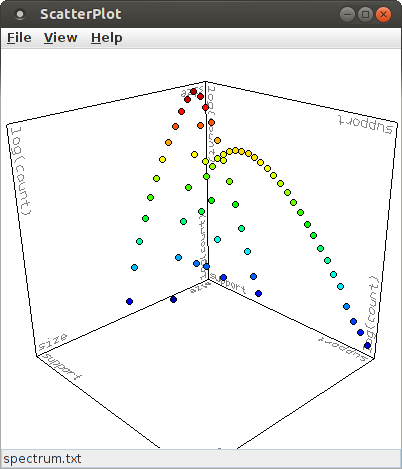
It is recommended to keep the selection for the X- and Y-axis at the pattern size and pattern support and to choose either the (average) occurrence count or its logarithm for the Z-axis. This yields an intuitive view of the mapping from pattern signatures to occurrence counts that a pattern spectrum represents.
The attribute chosen as an "Overlay" is used to determine the color of the points. By default, a rainbow color scale is used for all attributes. The color scale may be changed on a per attribute basis with the color selection dialog (see below).
The layout of the scatter plot can be changed with the dialog box that is opened with "View > Set Layout ...".
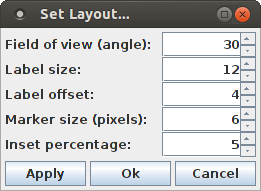
The layout dialog box allows to change the field of view of the projection, which is stated as an angle (namely the angle at which the viewing frustum opens at the eye point). The 3D projection is generally a central projection, which is the closer to a parallel projection, the lower the angle that defines the field of view.
The label size specifies the size of the labels that are written on cube faces, in pixels in a parallel projection. (Note that, as a consequence of the central projection, the label height may be greater or smaller than the number of pixels specified in the dialog box). The label offset states how many pixels the base line for the label text is away from the cube edges. The inset percentage states what percentage of the cube width the most extreme points are away from the cube faces. The default of 5% is usually a decent choice.
The colors used for the points in the scatter plot can be changed with the color dialog box that is opened with "View > Set Color ...", which is similar to the corresponding dialog box for a bar chart.
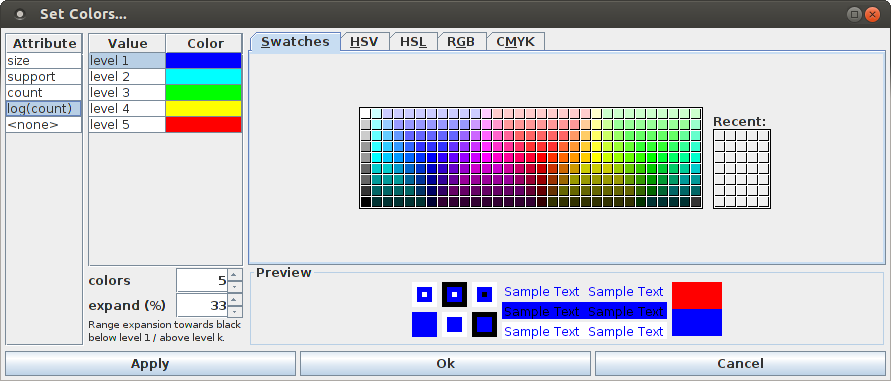
The right part of the color dialog box is a standard Java color selector, as it is also used for selecting the color of the dots in the dot display used to visualize parallel point processes/spike trains, or the dialog box used to specify the color scheme for the bars of a bar chart (see above). The left part of the dialog box contains a list of the attributes than can be chosen as a color overlay of the scatter plot, as the color selections that can be made for the scatter plot are attribute specific. The special attribute "<none>" serves the purpose to select the color of the points if no overlay is desired.
The middle part of the dialog box depends on the type of the attribute that is selected from the list on the left. If this attribute is nominal (that is, has only a finite set of possible values, as it is the case for the pattern size and support, which are treated as nominal attributes, or the special attribute "<none>", which has only one value), the middle part contains a simple list of these values. For each value a color can be chosen. By default, these values are chosen equidistant from a rainbow color scale with as many entries as there are values.
If the attribute selected in the list on the left is metric (numeric), like the (average) occurrence count or its logarithm, the middle part looks like the left part in the color selector box for the bar colors. The default is a list of five color levels, ranging from blue to red, with turquoise, green and yellow in the middle. The number of colors can be changed with the "colors" spinner below the color list, for example, it can be reduced to two to obtain a simple heat map scale. The second spinner, labeled "expand (%)" specifies by how much the color range is extended towards black below the color for level 1 and above the color for level k, where the percentage refers to the step between to colors (100%). An expansion factor of one third (33%) usually yields a visually pleasing color scale.
The points of the scatter plot are colored according to the attribute that is chosen as "Overlay" in the attributes selection dialog box. As already mentioned, "<none>" refers to no attribute, which means that the same color will be used for all points.
| back to the top |  |
![]()
The "View" button next to the label "Pattern output file" on the "Files" tab opens a simple table viewer for the patterns that were found in the original data (that is, the data as it is read from the (spike) trains file) and which survived the pattern spectrum filtering (PSF) and pattern set reduction (PSR) steps. Note that a found patterns are only available after the analysis process has been executed by pressing the "Run" button of the main CoCoGUI window and waiting for completion.
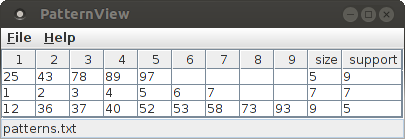
The found patterns are displayed in a simple table, which has as many columns as the largest pattern (one column per item/neuron) plus two columns that state the size and the support of the pattern.
| back to the top |  |
![]()
The following sections describe the data format of the three file types that the CoCoGUI program reads or writes: the (spike) trains file, the pattern spectrum file and the pattern output file.
| back to the top |  |
![]()
The (spike) trains file has to be a (text) file structured by field and record separators and blanks. Record separators, not surprisingly, separate records, usually lines (since the default record separator is the newline character), field separators separate fields, usually words (since among the default field separators are the space and the tabulator, but also the comma). Blanks are used to fill fields, for example, to align them. In addition, comment characters are recognized. If a record starts with a character that is among those declared as comment characters, the record is considered to be a comment and therefore ignored.
There are four different record formats, which can be selected on the "Format" tab, on which the record and field separators and the blank and comment characters can be specified. The four record formats are:
Note that in all of the above formats, commas or tabulator characters may also be used as the field separator without having to change any settings, since the comma and the tabulator character are also among the default field separators.
| back to the top |  |
![]()
A pattern spectrum maps pattern signatures, that is, a pair of the size of a pattern (number of items/neurons) and its support (number of (co-)occurrences in the maximum independent set sense) to occurrence counts, that is, to the (average) number of patterns with such a signature that have been observed in the surrogate data sets.
The pattern spectrum file contains three columns: the first column
contains the pattern size, the second column the pattern support and
the third column the (average) occurrence count. An example pattern
spectrum file starts like this:
2 2 27.35
2 3 82.748
2 4 188.141
2 5 338.811
2 6 504.222
2 7 632.721
2 8 693.565
2 9 666.21
2 10 574.587
2 11 445.557
2 12 316.625
...
By default the fields in this file are separated by spaces and the records are separated by newline characters (making them the lines of a simple text file). The field separator, which is a space by default, may be changed on the "Output" tab, but it is not recommended to do so as other field separators than a space, a tabulator character or a comma may make it impossible to open the file in the pattern spectrum viewer (see above).
The size and the support are always written as integer numbers, the (average) occurrence count as an integer or a floating point number. Note that a pattern spectrum file that has been created outside of the CoCoGUI program and that states the size and/or the support as floating point numbers may cause problems with the pattern spectrum viewer. The CoCoGUI program writes the triplets of (size, support, count) sorted by size, and within the same size sorted by support.
| back to the top |  |
![]()
The pattern output file lists the found patterns (after pattern spectrum filtering and pattern set reduction), one pattern per record. The records of this file are separated by newline characters, making them the rows of a simple text file. Each record contains one found pattern, described as a list of items/neurons and a support indicator. The items/neurons are separated by field separators, which is a space by default. The field separator can be changed on the "Output" tab, but it is not recommended to do so as other field separators than a space, a tabulator character or a comma may make it impossible to open the file in the pattern set viewer.
Furthermore, each record may be started with a fixed string, which is the empty string by default, and the format for the support value, which is printed after the list of items may be changed from its default " (%d)" (that is, the support is printed separated by a space and enclosed in parentheses as a decimal integer number). However, it is not recommended to do so as header strings and different support value formats may make it impossible to open the pattern output file with the pattern set viewer.
| back to the top |  |
![]()
| back to the top |  |
![]()
(MIT license, or more precisely Expat License; to be found in the file mit-license.txt in the directory coconad/doc in the source package, see also opensource.org and wikipedia.org)
© 2013-2016 Christian Borgelt
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
| back to the top |  |
![]()
Download page with most recent version.
| back to the top |  |
![]()
| E-mail: |
christian@borgelt.net christian.borgelt@softcomputing.es | |
| Snail mail: |
Christian Borgelt Intelligent Data Analysis and Graphical Models Research Unit European Center for Soft Computing Edificio Cientifico-Tecnológico, 3a Planta c/ Gonzalo Gutiérrez Quirós s/n 33600 Mieres (Asturias) Spain | |
| Phone: | +34 985 456545 | |
| Fax: | +34 985 456699 |
| back to the top |  |
![]()